About this course booklet
The proton motive force (Δp = ΔΨ_m − (2.303 RT/F)·ΔpH) is the central energetic currency of life. Generated by the electron transport chain, it captures a large fraction of the free energy released by NADH oxidation and redistributes it across ATP synthesis, transport processes, and signaling pathways.
Under physiological conditions (ΔΨ_m ≈ −150 to −180 mV, ΔpH ≈ 0.5–1), mitochondria convert redox energy into an electrochemical gradient of ~180–220 mV, corresponding to ~17–21 kJ/mol per proton.
This 13-page course booklet, authored in April 2026, provides a compact yet rigorous framework for understanding how Δp is generated, quantified, and perturbed in disease. It connects thermodynamic principles to real biological systems, with explicit numerical derivations and clinical implications.
Inside the booklet (5 modules)
Module 1 — What Is the Proton Motive Force?
- Mitchell (1961): the immediate product of respiration is not a chemical intermediate but an electrochemical field across the IMM.
- Δp has two additive components: electrical (ΔΨ_m) + voltage-equivalent chemical (ΔpH).
- Healthy Δp ≈ 180–220 mV → ~19–21 kJ/mol per proton.
Module 2 — The Electrical Component (ΔΨ_m)
- ETC is a vectorial charge pump. Matrix loses positive charge → electronegative (−150 to −180 mV, matrix-negative).
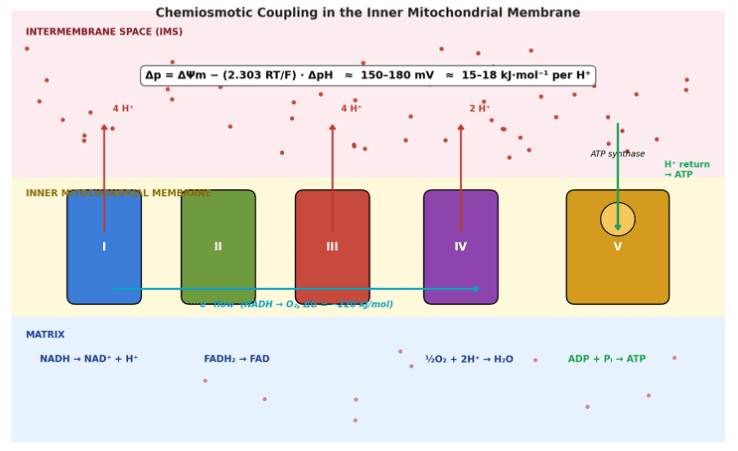
- Stoichiometry table: Complex I = 4 H⁺, III = 4 H⁺, IV = 2 H⁺ per 2 e⁻ (Complex II = 0).
- ~75–80% of total pmf resides in ΔΨ_m in mammalian mitochondria.
- Clinical: TMRM/JC-1 probes; metformin (Complex I), rotenone, oligomycin, FCCP; persistent depolarisation predicts mortality in sepsis.
Module 3 — The Chemical Component (ΔpH)
- Matrix alkaline by 0.5–1 pH unit due to proton ejection + matrix buffering (phosphate, carboxylates, histidines).
- ΔΨ_m and ΔpH are thermodynamically interconvertible: valinomycin collapses ΔΨ_m, nigericin collapses ΔpH, FCCP/CCCP/DNP collapse both.
- Jagendorf’s acid-bath experiment (1966, chloroplasts) proved ATP synthesis can run on ΔpH alone.
Module 4 — Quantifying the PMF (Nernst Factor)
- Full derivation: from Δμ̃_H+ = F·ΔΨ_m − 2.303 RT·ΔpH to the voltage form Δp = ΔΨ_m − (2.303 RT/F)·ΔpH.
- At 37 °C: 2.303 RT/F ≈ 61.5 mV/pH unit.
- Worked example: ΔΨ_m = −170 mV, ΔpH = 0.6 → Δp ≈ 207 mV → 20 kJ/mol H⁺ → 10 H⁺ × 20 = 200 kJ/mol captured per NADH → ~91 % of the 220 kJ/mol liberated by NADH → O2. At lower PMF (155–180 mV), the captured fraction drops to the textbook 150–175 kJ/mol (68–80 %) range.
- ATP synthesis ceiling: ΔG_ATP ≤ ~4 H⁺ × 20 kJ = 80 kJ/mol ATP; measured ~55–60 kJ/mol (~70 % of max). The shortfall is the flux-driving dissipation required by the second law.
Module 5 — Integration, Physiology & Clinical Relevance
- Δp governs: MCU-driven Ca²⁺ uptake, ROS production (reverse electron transport at Complex I), PINK1/Parkin mitophagy signalling, TIM23-dependent protein import.
- Controlled uncoupling (UCP1–3, therapeutic uncouplers like BAM15) dissipates Δp as heat — feature, not bug.
- Immunometabolic phenotypes correlate with Δp set-point: effector T cell / M1 macrophage = lowered, glycolytic, Warburg-like; memory / Treg = high ΔΨ_m, OXPHOS + FAO; senescent = low, unstable.
- Pmf-collapse cascade: ETC injury → depolarisation → ATP synthase reversal → MPTP → cyt c release → DAMP-driven sterile inflammation (cGAS-STING, TLR9).
- Therapeutic axes: NAD⁺ precursors (NMN, NR); cardiolipin stabilisation (SS-31/elamipretide); TPP⁺-targeted drugs (MitoQ, MitoTEMPO); mitochondrial transplantation.
Key equations
- PMF (Mitchell): Δp = ΔΨ_m − (2.303 RT/F)·ΔpH
- Energy per proton: ΔG = F·Δp ≈ 17–21 kJ/mol
- Captured per NADH: 10 H⁺ × F·Δp ≈ 150–200 kJ/mol
- Total redox drop NADH → O₂: ΔG°’ = −n·F·ΔE°’ = −2·96485·1.14 V ≈ −220 kJ/mol
- Nernst factor (37 °C): 61.5 mV / pH unit
- ATP synthesis ceiling: ΔG_ATP ≤ n·F·Δp ≈ 80 kJ/mol (measured −55 to −60 kJ/mol)
Who is this for?
- PhD students in life sciences, biophysics, and bioenergetics
- Medical doctors seeking a quantitative understanding of mitochondrial function
- Researchers in redox biology and systems metabolism
- Clinicians exploring immunometabolism and mitochondrial medicine
Key insight
The proton motive force is the universal electrochemical battery of life: generated by respiration, consumed by cellular work, and whose collapse is a quantitative signature of disease.
Note: The full PDF is in English.